Treangen Lab, in Rice University’s Department of Computer Science, is the recipient of a contract from the Centers for Disease Control and Prevention (CDC) to develop the Harvest Variant software for integrated, collaborative real-time variant tracking of SARS-CoV-2. The ability to conduct real-time monitoring will support both CDC and the broader public health community’s understanding of changes to SARS-CoV-2 infectivity and transmissibility, vaccine efficacy, and fitness within human hosts.
Todd Treangen, assistant professor of computer science, says, “It’s very exciting that my group will be providing robust software solutions specific to SARS-CoV-2 variant analyses and pathogen surveillance in support of CDC’s mission.”
The 12-month contract, “Harvest Variants: Enhancing Harvest tools for integrated, real-time, collaborative variant tracking of SARS-CoV-2,” includes a $630,000 grant and is a part of CDC’s SARS-CoV-2 Sequencing for Public Health Emergency Response, Epidemiology, and Surveillance (SPHERES) Initiative. These awards are intended to fill knowledge gaps and promote innovation in the U.S. in response to the COVID-19 pandemic. Funding awards are determined through a competitive selection process.
“The overarching goal is to leverage my existing software platform for microbial genome analyses to provide a cohesive, comprehensive software toolkit to facilitate analysis of SARS-CoV-2 variants by scientists in public health labs across the U.S.,” said Treangen.
With this new software upgrade, SARS-CoV-2 researchers will be able to track variants of interest and the variants of concern, such as the Delta variant. Further development of the Harvest code base will help regional scientists identify and track variants’ progression, based on location, infectivity rates or other factors. Since the onset of the COVID-19 pandemic, there have been several variants of note in the spike glycoprotein that have been linked to increased infectivity and are under active investigation.
The software was originally jointly developed with NIH scientists Sergey Koren, Brian Ondov, and Adam Phillippy and was published in 2014. Since publication, it has been cited nearly a thousand times. The software suite has three parts, including Parsnp, Harvest tools and Gingr, an interactive, graphic user interface.
In addition to Treangen, lab members participating on the project are software engineer Matt Barnett as well as doctoral students Bryce Kille and Yunxi Liu.
When discussing the new CDC grant, Kille says, “I’m excited! I really enjoy the fast pace of such short production timelines.” His goal is to “make sure the software development process goes smoothly and builds are delivered on time.”
Barnett adds, “I can think of few ways better to meet our institutional charge than by working with the CDC to help create the knowledge needed to combat perhaps the principal acute public health crisis of our time.”
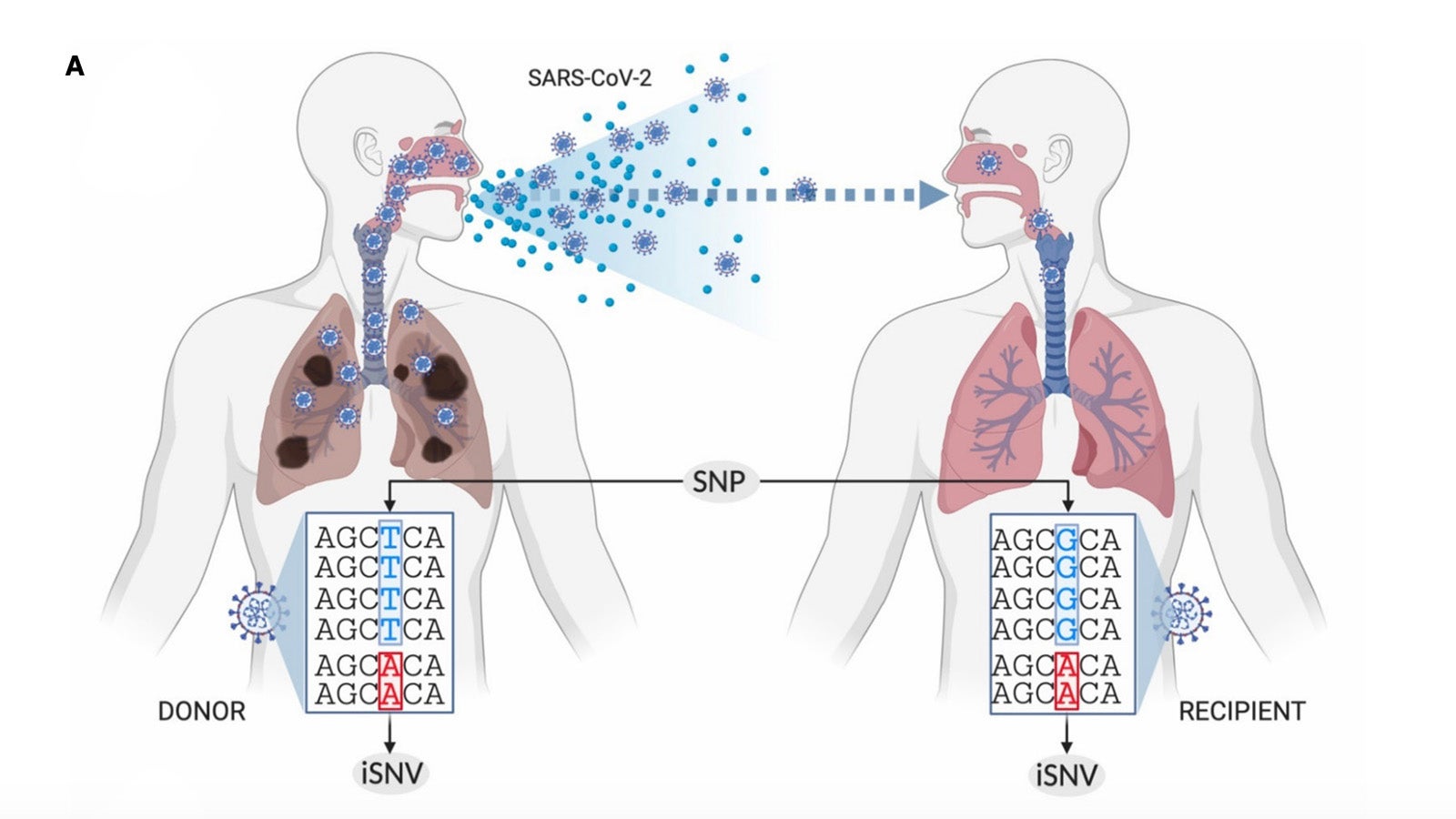
Real-time monitoring of viral mutations of circulating SARS-CoV-2 lineages, both at the intrahost and interhost level, is a key step in understanding changes to SARS-CoV-2 infectivity/transmissibility, vaccine efficacy, and fitness within human hosts. The Treangen Lab will focus on the design, implementation, optimization, and deployment of the Harvest Variants software. Signature Science, LLC will support this project by providing functional SARS-CoV-2 biocurations and annotations of variants of concern. This work will help end users visualize and interpret the significance of observed SARS-CoV-2 mutations.